Research
Unlocking Protein Dynamics
Biology is a study of motion and change — yet we still rely on frozen snapshots to understand it.
Static structures can’t tell us what makes a designed enzyme functional, how dynamics shape drug efficacy and resistance, or how motion itself can be engineered and optimized. These are the questions at the frontier of drug discovery, enzyme design, and disease research.
The DiffUSE project is building the methods, tools, and infrastructure to make protein motion visible, measurable, and usable, so this frontier becomes accessible to endless biological questions.
Four Areas of Focus
Methods
We are supporting ambitious technical moonshots to improve how dynamic structural biology data is collected, processed, and extracted, with the goal of democratizing these methods so that capturing protein motion becomes routine.
Modeling
We are developing machine learning algorithms to uncover hidden information in experimental data, build more robust ensemble models, and leverage information across multiple experimental sources, enabling improved future ensemble prediction models.
Infrastructure
We are establishing standards and infrastructure needed to deposit, validate, search, and prepare dynamic structural data for machine learning at scale. Make dynamic data as accessible as static structures so that anyone, not just structural biologists, can leverage it to answer biological questions.
Biological Impact
We are developing tools to elucidate how conformational ensembles drive function, from binding specificity and catalysis to mutational effects, by building the computational tools and metrics that make ensemble-aware enzyme design a practical reality.
Our First Experimental Deep Dive: Diffuse Scattering
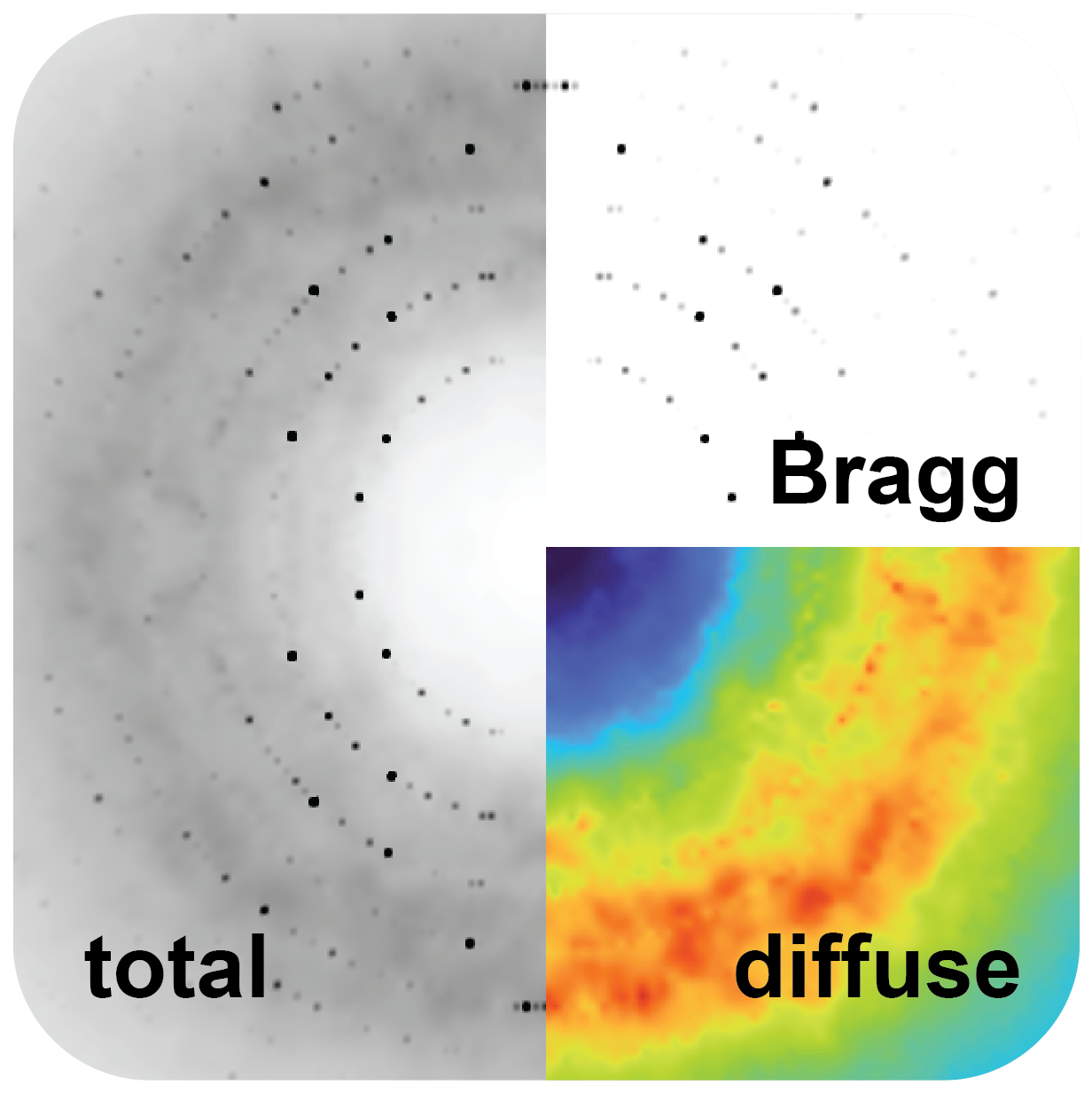
Our first experimental deep dive is expanding the use of diffuse scattering: a largely overlooked signal measured by X-ray crystallography that could unlock our ability to measure protein dynamics.
-
The DiffUSE Project Logbook: Catalog X-ray datasets, collect notes from synchrotron data collection, share reports from data processing, and post preliminary analysis without filters.
-
The DiffUSE Data Management App: Under active development, this site will function to document experiment processing with markdown, images, tables, and rich metadata fields for comprehensive research tracking across datasets and beamlines.
-
Posts from our Scientists: Read direct dispatches from our distributed DiffUSE team.
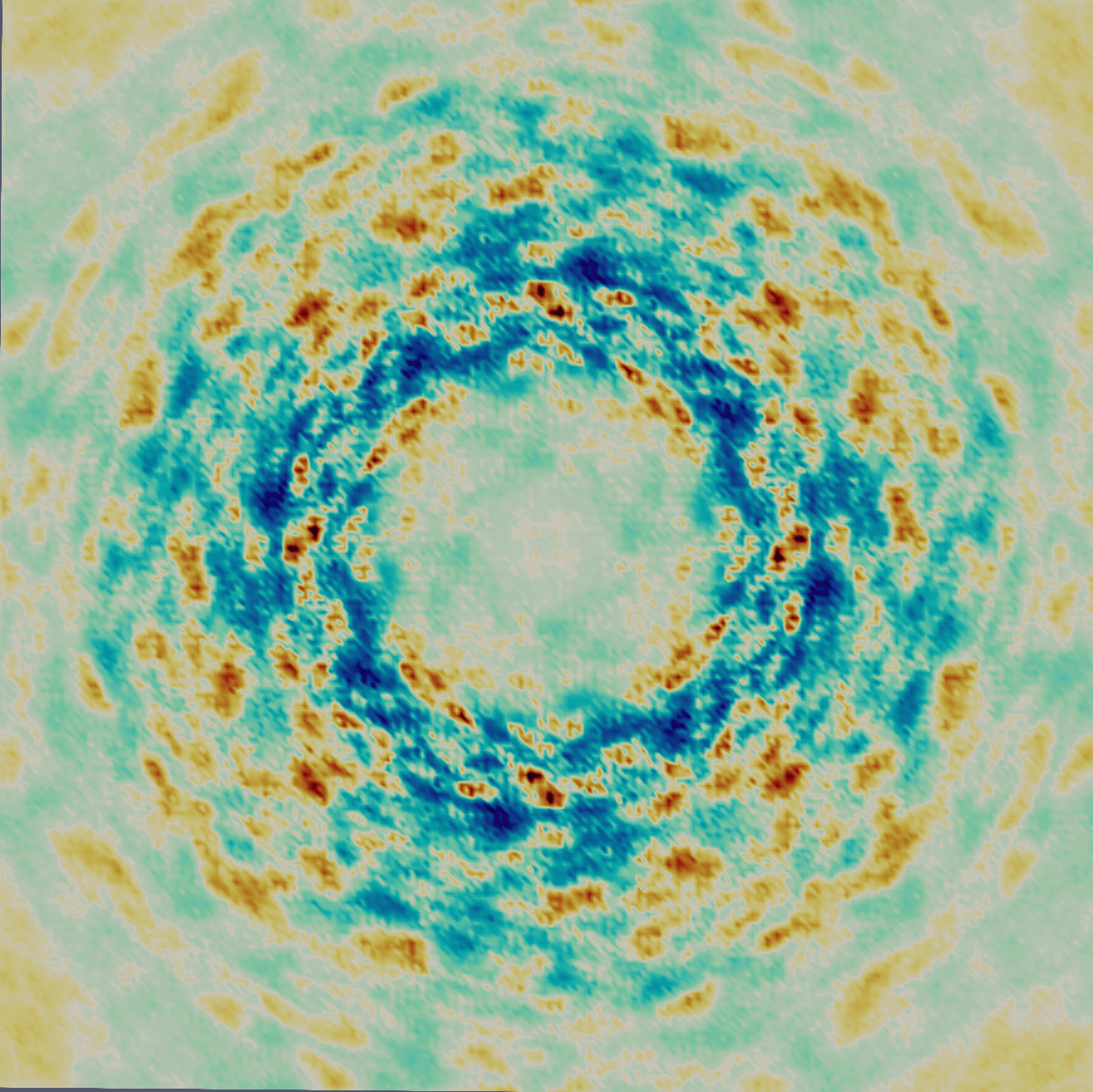
Enable data collection

Most crystallography beamlines are optimized for collecting bright diffraction signals from cryogenically frozen samples to simplify transport and reduce radiation damage. To study protein dynamics with the much weaker and complex diffuse scattering signals, we need unfrozen samples, which introduces new challenges. We are developing practical strategies for measuring diffuse scattering at room temperature, addressing experimental constraints to make this accessible for non-specialists.
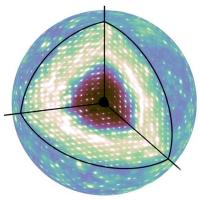
Make software accessible
Until now, diffuse scattering analysis has been carried out by a small number of experts using custom-built software. We will build on the state of the art to develop tools that are intuitive, robust, and usable by the broader community.
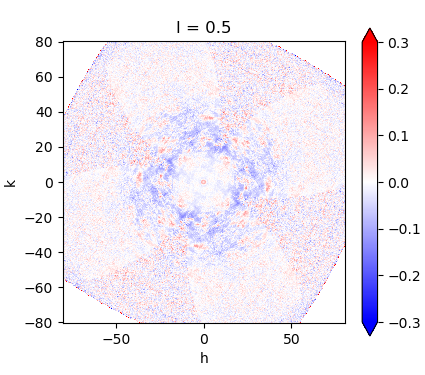
Expand public datasets
With only a handful of publicly available diffuse scattering datasets today, we will lead an effort to populate a public database with compelling, high-quality examples, expanding the resources available to the community and lowering barriers for others to analyze, interpret, and apply diffuse scattering data in their research.
Improve models of protein dynamics
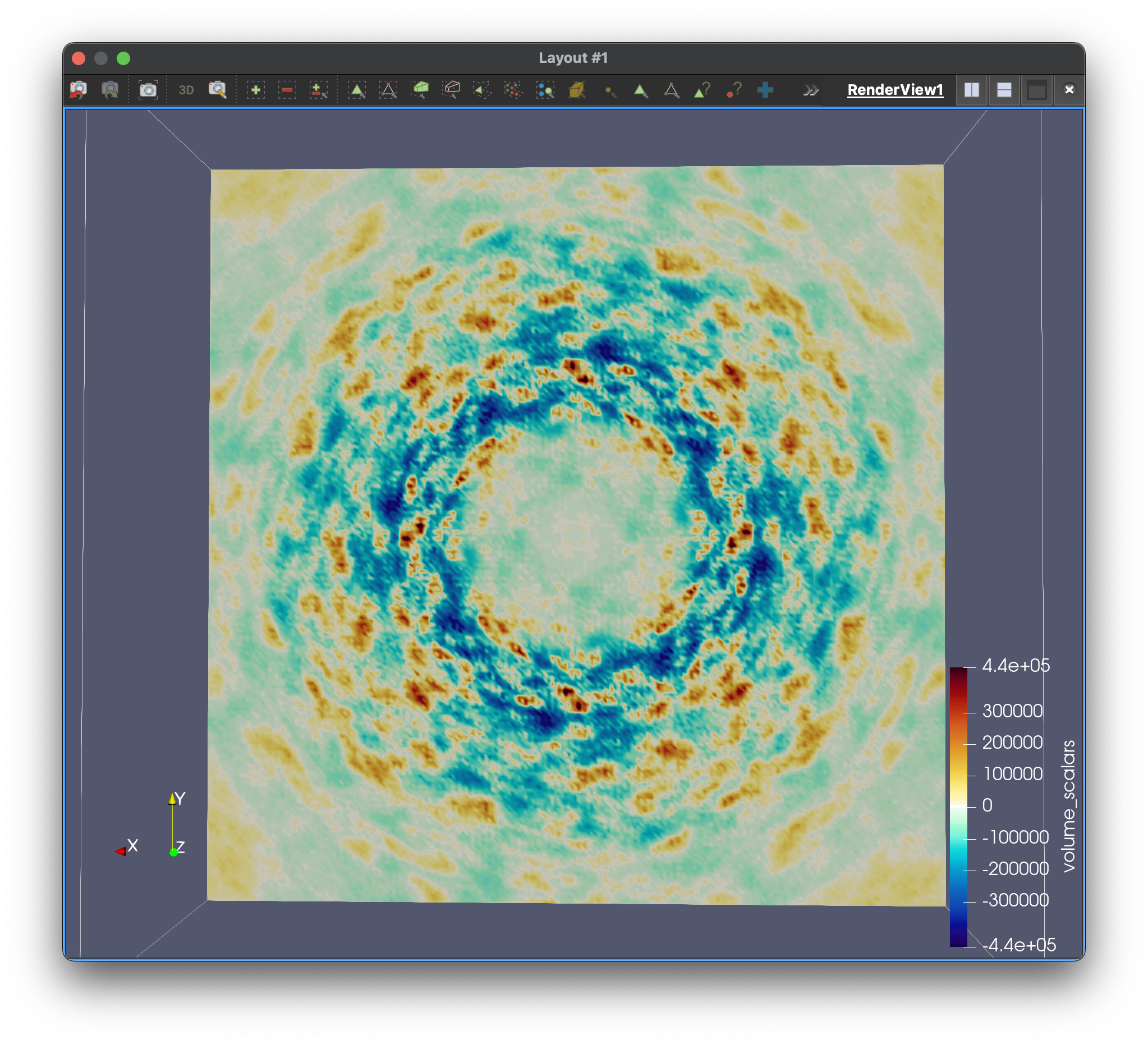
To extract meaningful biological insights from diffuse scattering data, accurate modeling is essential. At present, we model protein motions separately from Bragg peaks and from diffuse scattering, then compare them, but the results often disagree. We are developing algorithms that learn directly from experimental data, integrating information from both Bragg and diffuse scattering to develop a unified, consistent model.
Encoding and validating dynamic models
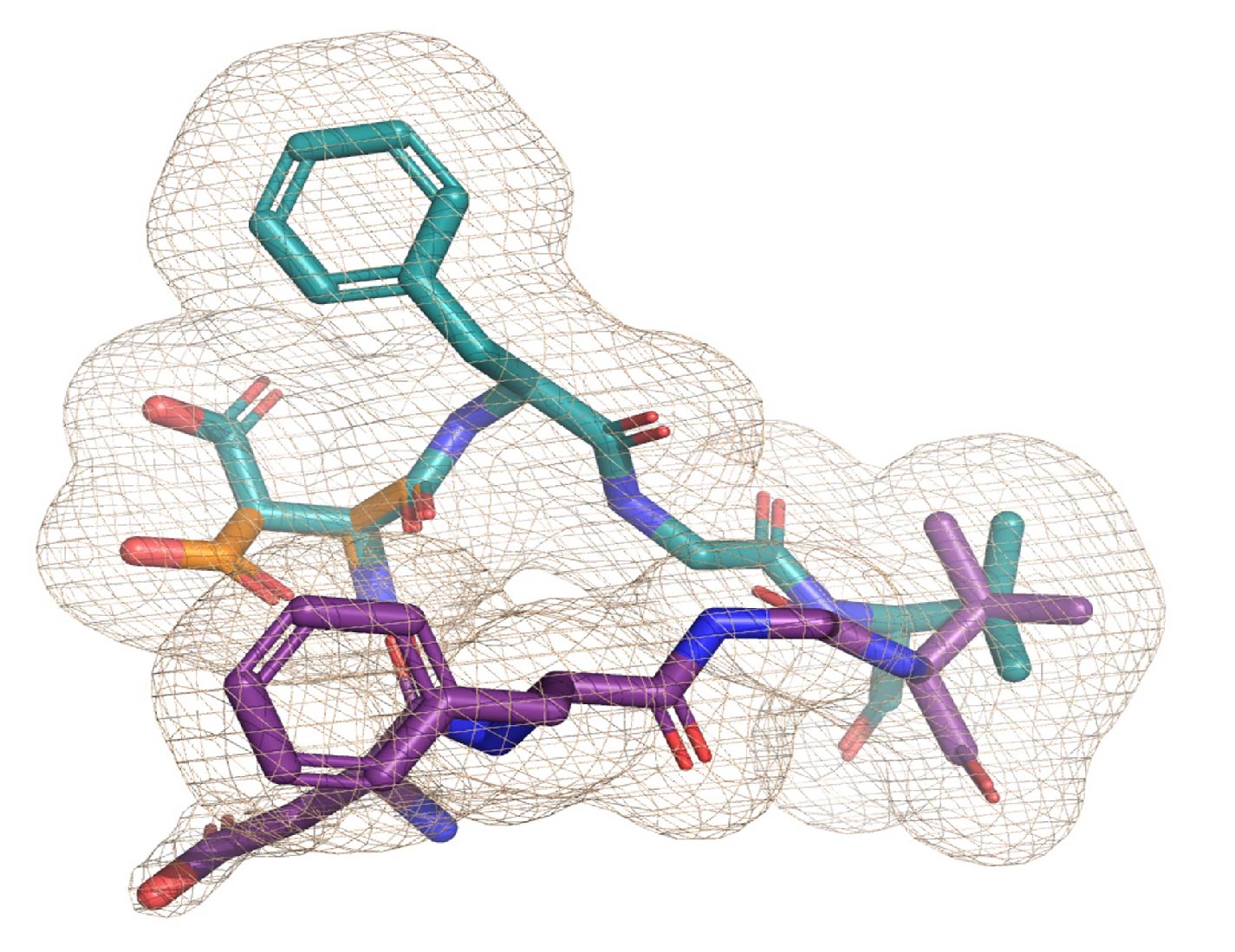
Correctly representing and encoding the outputs of structural modeling is essential for translating them into biological insight. To enable AI co-driven scientific discovery, these representations must be both machine-readable and human-interpretable to leverage the full richness of existing and future experimental data. We are developing infrastructure to improve model encoding and validation, enabling others to learn from the rich experimental data collected in this project.